perEditor
A tool to create personalized genome sequences
Home Download Installation Usage VCF format Tutorial
Tutorial
To help you understand how perEditor and perEditor_ra work you can try the following examples.
perEditor
chr1.fa: reference sequence in fasta format.
variations.vcf:
SNP/indels annotation VCF file. This file contains
information about 629 individuals. For each individual the VCF file
specifies if the variations is on the maternal or paternal chromosome.
Go to the folder where you have downloaded these files and type:
perEditor
chr1.fa variations.vcf mother 1 chr1_customized.fa
This will tell to perEditor
that chr1.fa is the reference sequence that is going to be
customized using the information contained in variations.vcf. As variations.vcf contain information about several individuals (629) we are specifying that we want a customized reference relative to the maternal
allele (mother) of the first individual (1). The customized reference sequence will be
stored as chr1_customized.fa.
perEditor_ra
Download a set of sample input files from here. These sample files include four chromosomes
(fasta files), a rearrangement file (VCF 4.1 format), and two BED files with
information about the length of each chromosome and the location of its
centromeres, respectively. The rearrangement file specifies the following chromosomal modifications:
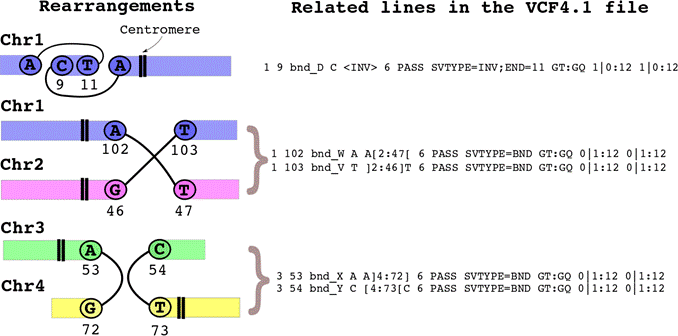
Once you have downloaded these files in a folder, go to that folder and type:
perEditor_ra
rearrangements_t.vcf chr_length_t.bed centromeres_t.bed 1 father
This will tell perEditor_ra that you are interested in the
parternal (father) allele of the first (1) individual. After do
so, perEditor_ra will create four new
chromosomes, namely chr1_new.fa, chr2_new.fa, chr3_new.fa, and chr4_new.fa that
have been modified to show the rearrangement contained in the rearrangements_t.vcf file.